Check out our Github space for code and programs
https://github.com/koszullab/
https://github.com/koszullab/
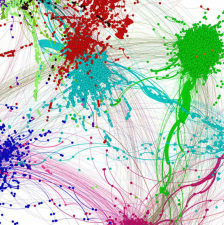
META3C (MetaHiC)
We have pushed further our analysis about the potential of 3D contact data to solve genomics limitations and problems. We have developed meta3C, a metagenomic chromosome conformation capture approach that allows characterizing individual genomes and their average chromosome organization within a mix of organisms. Meta3C can be applied to species already sequenced, or can be used directly for de novo assembling, scaffolding and characterizing the tridimensional organization of unknown genomes.
See Marbouty et al., 2014 (b)
We have pushed further our analysis about the potential of 3D contact data to solve genomics limitations and problems. We have developed meta3C, a metagenomic chromosome conformation capture approach that allows characterizing individual genomes and their average chromosome organization within a mix of organisms. Meta3C can be applied to species already sequenced, or can be used directly for de novo assembling, scaffolding and characterizing the tridimensional organization of unknown genomes.
See Marbouty et al., 2014 (b)
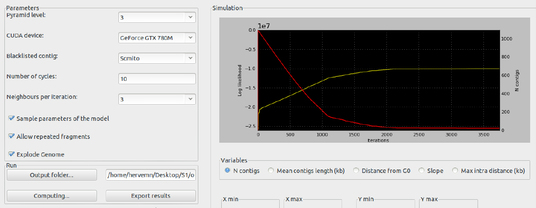
GRAAL (Genome (Re)Assembly Assessing Likelihood from 3D)
GRAAL generates high-quality assemblies of genomes containing large repeated regions, and offers a direct probabilistic interpretation of these structures. GRAAL is a powerful and elegant method described in Marie-Nelly et al. (2014) that uses 3D contacts of DNA in nuclear space to efficiently finalize genome assemblies. In contrast to standard assembly techniques, our method accurately recovers the actual number and DNA content of chromosomes, and does not require expensive and lengthy follow-up experiments.
GRAAL readme file (pdf)
GRAAL program (zip)
See Marie-Nelly et al., 2014 (b)
GRAAL generates high-quality assemblies of genomes containing large repeated regions, and offers a direct probabilistic interpretation of these structures. GRAAL is a powerful and elegant method described in Marie-Nelly et al. (2014) that uses 3D contacts of DNA in nuclear space to efficiently finalize genome assemblies. In contrast to standard assembly techniques, our method accurately recovers the actual number and DNA content of chromosomes, and does not require expensive and lengthy follow-up experiments.
GRAAL readme file (pdf)
GRAAL program (zip)
See Marie-Nelly et al., 2014 (b)
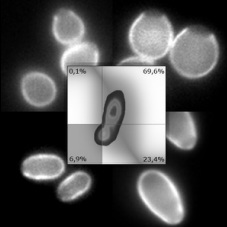
ELUTRIATION OF YEAST SPECIES
Centrifugal elutriation discriminates cells according to their sedimentation coefficient, generating homogeneous samples well suited for genomic comparative approaches. It can, for instance, isolate G1 daughter cells from a Saccharomyces cerevisiae unsynchronized population, alleviating aging and cell-cycle biases when conducting genome-wide/single-cells studies. We propose a straightforward and robust procedure to determine whether a cell population of virtually any yeast species can be efficiently elutriated, while offering solutions to optimize success. This approach was used to characterize elutriation parameters and S-phase progression of four yeast species (S. cerevisiae, Candida glabrata, Lachancea kluyveri and Pichia sorbitophila) and could theoretically be applied to any culture of single, individual cells.
See Marbouty et al., 2014 (a)
Centrifugal elutriation discriminates cells according to their sedimentation coefficient, generating homogeneous samples well suited for genomic comparative approaches. It can, for instance, isolate G1 daughter cells from a Saccharomyces cerevisiae unsynchronized population, alleviating aging and cell-cycle biases when conducting genome-wide/single-cells studies. We propose a straightforward and robust procedure to determine whether a cell population of virtually any yeast species can be efficiently elutriated, while offering solutions to optimize success. This approach was used to characterize elutriation parameters and S-phase progression of four yeast species (S. cerevisiae, Candida glabrata, Lachancea kluyveri and Pichia sorbitophila) and could theoretically be applied to any culture of single, individual cells.
See Marbouty et al., 2014 (a)
